|
Back to Blog
Cytoscape 2.86/17/2023 Model trees are very useful techniques to estimate the numerical values for the target genes by linear regression functions. REGNET generates gene association networks from gene expression data, and differs from correlation-based methods in that the relationship between one gene and others is calculated simultaneously. This experiment shows that REGNET performs more accurately at detecting true gene associations than the Pearson and Spearman zeroth and first-order correlation-based methods. 
Second the E.coli transcriptional network (in the Regulon database) is used as control to compare the results to that of a correlation-based method. First, the biological coherence of the results are tested. The performance of our approach, named REGNET, is experimentally tested on two well-known data sets: Saccharomyces Cerevisiae and E.coli data set. For this reason we apply a statistical procedure to control the false discovery rate. Finally, a graph from all the relationships among output and input genes is built taking into account whether the pair of genes is statistically significant. While correlation-based methods analyze each pair of genes, in our approach we generate a single regression tree for each gene from the remaining genes. We propose model trees as a method to identify gene interaction networks. Correlation-based methods are very useful in order to determine whether two genes have a strong global similarity but do not detect local similarities.Results They are typically generated using correlation statistics as pairwise similarity measures. To infer gene regulatory networks, the first step is to find direct regulatory relationships between genes building the so-called gene co-expression networks. Novel strategies are required in order to handle the huge amount of data produced by microarray technologies. All core developers of Cytoscape are available to answer your questions.RegNetC: Inferring Regression Networks as a plugin for Cytoscape 2.8.2 To try the new version, please visit here. PSICQUIC Client in Cytoscape 3Ĭytoscape 3.1.0 has been released since 2014 and the PSICQUIC client feature is a built-in function from that version. For more information about MIQL, please read the reference page. You can simply type the query in the text field. If you press the Merge button, the Advanced Network Marge plugin will be started and you can merge imported networks manually. When the import process is finished, you also have the option to edit the network titles if you want. If the result is 0 or the service is inactive, it will be ignored. After the search process, it pops up a window, where you can select which data sets you want to import. You can simply copy and paste list of identifiers to the text box. In this mode, Cytoscape sends list of identifiers as query. You can switch the mode from Search Property tab. There are two search modes in the client: If PSICQUIC client is properly installed, you can find it in Data Source combo box. To start the plugin, select the client from File->Import->Network from web services. Select Plugin->Manage Plugins and then you can find the plugin under Network and Attribute IO section -> PSICQUICUniversalClient Start Plugin The Plugin is available from Cytoscape Plugin Manager. If you have older versions, please download the latest version. The PSICQUIC Client Plugin is compatible with Cytoscape 2.7.0 or later. How to Use PSICQUIC Client in Cytoscape 2 Install Plugin From a single window, you can send your query to all of registered PSICQUIC services at once. You can access PSICQUIC databases just like the other clients. PSICQUIC Client for Cytoscape is a part of Cytoscape's web service client framework. For more information about Cytoscape, please visit official web site. The advantage of using PSICQUIC services with Cytoscape client is you can save your work as a snapshot (session file) and also you can use its powerful visualization features. PSICQUIC Universal Client is a Cytoscape plugin for querying multiple PSICQUIC-compliant interaction data services from a simple user interface. It is widely used in systems biology community and is expandable by plugins. This is compatible with Cytoscape 2.7.0 or later.Ĭytoscape is an open source platform for general network data integration, analysis, and visualization. : Now PSICQUIC Client is part of Cytoscape 3 core!.Bug fixes for edge score import function.Smart Label - Plugin adds more human-readable gene names to the node. 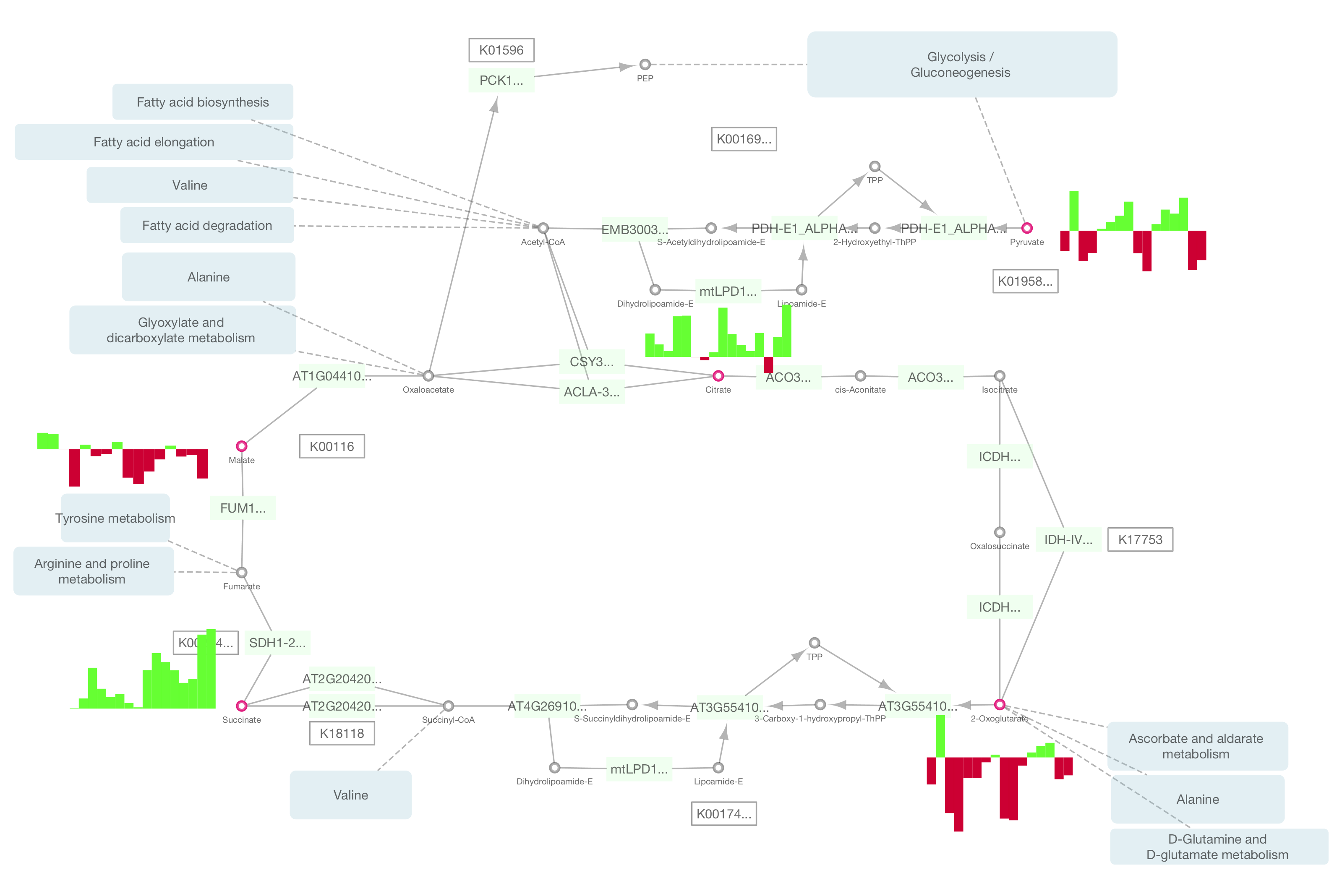
Please use with latest version of Cytoscape (2.8.3). This update is recommended for all current users.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
